Apostle MiniGenomics technology is a modified extension to our flagship Apostle MiniMax technology. Apostle MiniGenomics focuses on providing an efficient and economic solution for isolating and purifying genomic nucleic acids from a spectrum of biological medium, supporting the accurate subsequent genetic and genomic analyses.
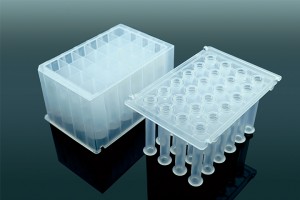
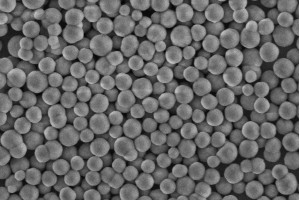
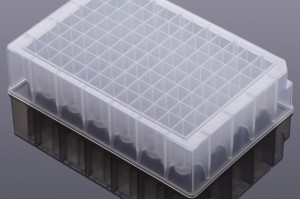
Empowered by the Apostle MiniGenomics technology, Apostle COVID-19 RNA Extraction System is composed of an Apostle MagTouch Nucleic Acids Isolation Automation System, an Apostle MiniGenomics Viral TotalNA Isolation Fast Kit, Apostle 96-Well Deep Well Plates and Tip Combs. The system is based on magnetic nanoparticle technologies designed for fast extraction and purification of viral nucleic acids from various kinds of biological samples collected in multiple transport media. To date, our clients have processed more than 20 million swabs in various CAP/CLIA clinical labs in the United States. It's also included in a US FDA EUA authorized SARS-CoV-2 molecular diagnostic test: Fulgent COVID-19 by RT-PCT Test.
In addition, Apostle MiniGenomics High Efficiency Urinary Tract Microbiota DNA Isolation Kit is designed for isolation of microbial DNA from urine samples. The kit offers highly efficient, reproducible recovery of high-quality bacterial DNA with high yield. The isolated DNA samples are suitable for a broad range of subsequent applications, including sequencing, PCR, etc.
Apostle MiniGenomics technology has also been used in Stool Sample Collection Kit, DNA Isolation Kit together with Sample Pretreatment Kit for Methylation Detection, for which our clients have successfully obtained CE mark.
Apostle MiniGenomics technology can be utilized in a broad spectrum of nucleic acid isolation applications from a range of biological medium, offering accuracy, efficiency, and low cost.