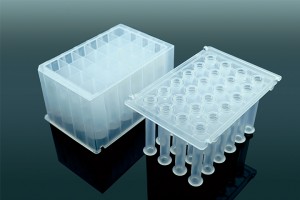
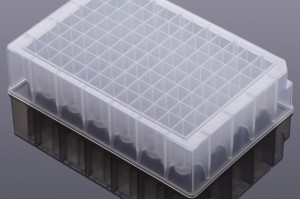
Apostle technologies have been applied in many world-class R&D studies, clinical laboratory settings, and public health response and surveillance.
This page lists some of the examples in Next-Gen Sequencing (NGS).
For a complete list of applications citing Apostle technologies, including publications and customer testimonials, see References.
Integrative modeling of tumor genomes and epigenomes for enhanced cancer diagnosis by cell-free DNA.
Mingyun Bae, Gyuhee Kim, Tae-Rim Lee, et al. Nature Communications 14, Article number: 2017 (2023)
(Note: Apostle MiniMax technology is used in this study.)
Abstract Multi-cancer early detection remains a key challenge in cell-free DNA (cfDNA)-based liquid biopsy. Here, we perform cfDNA whole-genome sequencing to generate two test datasets covering 2125 patient samples of 9 cancer types and 1241 normal control samples, and also a reference dataset for background variant filtering based on 20,529 low-depth healthy samples. An external cfDNA dataset consisting of 208 cancer and 214 normal control samples is used for additional evaluation. Accuracy for cancer detection and tissue-of-origin localization is achieved using our algorithm, which incorporates cancer type-specific profiles of mutation distribution and chromatin organization in tumor tissues as model references. Our integrative model detects early-stage cancers, including those of pancreatic origin, with high sensitivity that is comparable to that of late-stage detection. Model interpretation reveals the contribution of cancer type-specific genomic and epigenomic features. Our methodologies may lay the groundwork for accurate cfDNA-based cancer diagnosis, especially at early stages..
(Methods section) cfDNA was extracted from 0.4 mL plasma ... and eluted in a final volume of 22 μL, using an Apostle MiniMax High Efficiency cfDNA Isolation Kit (Apostle, US) according to the manufacturer’s instructions.
Individualized, heterologous chimpanzee adenovirus and self-amplifying mRNA neoantigen vaccine for advanced metastatic solid tumors: phase 1 trial interim results.
Christine D. Palmer, Amy R. Rappaport, Matthew J. Davis, et al. Nature Medicine volume 28, pages 1619–1629 (2022)
Abstract Checkpoint inhibitor (CPI) therapies provide limited benefit to patients with tumors of low immune reactivity. T cell-inducing vaccines hold promise to exert long-lasting disease control in combination with CPI therapy. Safety, tolerability and recommended phase 2 dose (RP2D) of an individualized, heterologous chimpanzee adenovirus (ChAd68) and self-amplifying mRNA (samRNA)-based neoantigen vaccine in combination with nivolumab and ipilimumab were assessed as primary endpoints in an ongoing phase 1/2 study in patients with advanced metastatic solid tumors (NCT03639714). The individualized vaccine regimen was safe and well tolerated, with no dose-limiting toxicities. Treatment-related adverse events (TRAEs) >10% included pyrexia, fatigue, musculoskeletal and injection site pain and diarrhea. Serious TRAEs included one count each of pyrexia, duodenitis, increased transaminases and hyperthyroidism. The RP2D was 1012 viral particles (VP) ChAd68 and 30 µg samRNA. Secondary endpoints included immunogenicity, feasibility of manufacturing and overall survival (OS). Vaccine manufacturing was feasible, with vaccination inducing long-lasting neoantigen-specific CD8 T cell responses. Several patients with microsatellite-stable colorectal cancer (MSS-CRC) had improved OS. Exploratory biomarker analyses showed decreased circulating tumor DNA (ctDNA) in patients with prolonged OS. Although small study size limits statistical and translational analyses, the increased OS observed in MSS-CRC warrants further exploration in larger randomized studies.
(Methods section) cfDNA was extracted from the entire plasma volume of a single draw using the Apostle MiniMax cfDNA Isolation kit (ApostleBio)
(Methods section) Enriched libraries were sequenced on an Illumina NovaSeq to a minimum mean raw depth of 80,000... Variant allele frequency was converted to mutated haploid genomic equivalents per milliliter plasma (hGE ml1 ) using the extracted plasma volume and total cfDNA yield from the extraction. Percent change in ctDNA was calculated as the change of the median mutated hGE ml1 from the baseline sample.
Simultaneous analysis of mutations and methylations in circulating cell-free DNA for hepatocellular carcinoma detection.
Pei Wang, Qianqian Song, Jie Ren, Weilong Zhang, Yuting Wang, Lin Zhou, Dongmei Wang, Kun Chen, Liping Jiang, Bochao Zhang, Wanqing Chen, Chunfeng Qu, Hong Zhao, Yuchen Jiao. National Cancer Center of China. Science Translational Medicine 14, eabp8704 (2022) 23 November 2022
Abstract Cell-free DNA (cfDNA)–based liquid biopsy is a promising approach for the early detection of cancer. A major hurdle is the limited yield of cfDNA from one blood draw, limiting the use of most samples to one test of either mutation or methylation. Here, we develop a technology, Mutation Capsule Plus (MCP), which enables multiplex profiling of one cfDNA sample, including simultaneous detection of genetic and epigenetic alterations and genome-wide discovery of methylation markers. With this technology, we performed de novo screening of methylation markers on cfDNA samples from 30 hepatocellular carcinoma (HCC) cases and 30 non-HCC controls. The methylation markers enriched in HCC cfDNA were further profiled in parallel with a panel of mutations on a training cohort of 60 HCC and 60 non-HCC cases, resulting in an HCC detection model. We validated the model in an independent retrospective cohort with 58 HCC and 198 non-HCC cases and got 90% sensitivity with 94% specificity. Furthermore, we applied the model to a prospective cohort of 311 asymptomatic hepatitis B virus carriers with normal liver ultrasonography and serum AFP concentration. The model detected four of the five HCC cases in the cohort, showing 80% sensitivity and 94% specificity. These findings demonstrate that the MCP technology has potential for the discovery and validation of multiomics biomarkers for the noninvasive detection of cancer. This study also provides a comprehensive database of genetic and epigenetic alterations in the cfDNA of a large cohort of HCC cases and high-risk non-HCC individuals.
(Methods Section) cfDNA was extracted from the plasma samples using the Apostle MiniMax cfDNA isolation kit (C43468, Apostle).
Reliability of Cell-Free DNA and Targeted NGS in Predicting Chromosomal Abnormalities of Patients With Myeloid Neoplasms.
Andrew Ip et al. Frontiers in Oncology June 14, 2022; https://doi.org/10.3389/fonc.2022.923809
Introduction: Cytogenetic analysis is important for stratifying patients with various neoplasms. We explored the use of targeted next generation sequencing (NGS) in detecting chromosomal structural abnormalities or copy number variations (CNVs) in patients with myeloid neoplasms.
Methods: Plasma cell-free DNA (cfDNA) from 2821 myeloid or lymphoid neoplasm patients were collected. cfDNA was sequenced using a 275 gene panel. CNVkit software was used for analyzing and visualizing CNVs. Cytogenetic data from corresponding bone marrow (BM) samples was available on 89 myeloid samples.
Results: Of the 2821 samples, 1539 (54.5%) showed evidence of mutations consistent with the presence of neoplastic clones in circulation. Of these 1539 samples, 906 (59%) showed abnormalities associated with myeloid neoplasms and 633 (41%) with lymphoid neoplasms. Chromosomal structural abnormalities in cfDNA were detected in 146 (16%) myeloid samples and 76 (12%) lymphoid samples. Upon comparison of the myeloid samples with 89 BM patients, NGS testing was able to reliably detect chromosomal gain or loss, except for fusion abnormalities. When cytogenetic abnormalities were classified according to prognostic classes, there was a complete (100%) concordance between cfDNA NGS data and cytogenetic data.
Conclusions: This data shows that liquid biopsy using targeted NGS is reliable in detecting chromosomal structural abnormalities in myeloid neoplasms. In specific circumstances, targeted NGS may be reliable and efficient to provide adequate information without the need for BM biopsy considering broad mutation profiling can be obtained through adequate sequencing within the same test. Overall, this study supports the use of liquid biopsy for early diagnosis and monitoring of patients with myeloid neoplasms.
We extracted cfDNA from 2821 peripheral blood samples using the Apostle MiniMax High Efficiency cfDNA Isolation Kit (San Jose, CA).
Efficient detection and post-surgical monitoring of colon cancer with a multi-marker DNA methylation liquid biopsy.
Jin et al. PNAS February 2, 2021 118 (5) e2017421118; https://doi.org/10.1073/pnas.2017421118
Multiplex assays, involving the simultaneous use of multiple circulating tumor DNA (ctDNA) markers, can improve the performance of liquid biopsies so that they are highly predictive of cancer recurrence. We have developed a single-tube methylation-specific quantitative PCR assay (mqMSP) that uses 10 different methylation markers and is capable of quantitative analysis of plasma samples with as little as 0.05% tumor DNA. In a cohort of 179 plasma samples from colorectal cancer (CRC) patients, adenoma patients, and healthy controls, the sensitivity and specificity of the mqMSP assay were 84.9% and 83.3%, respectively. In a head-to-head comparative study, the mqMSP assay also performed better for detecting early-stage (stage I and II) and premalignant polyps than a published SEPT9 assay. In an independent longitudinal cohort of 182 plasma samples (preoperative, postoperative, and follow-up) from 82 CRC patients, the mqMSP assay detected ctDNA in 73 (89.0%) of the preoperative plasma samples. Postoperative detection of ctDNA (within 2 wk of surgery) identified 11 of the 20 recurrence patients and was associated with poorer recurrence-free survival (hazard ratio, 4.20; P = 0.0005). With subsequent longitudinal monitoring, 14 patients (70%) had detectable ctDNA before recurrence, with a median lead time of 8.0 mo earlier than seen with radiologic imaging. The mqMSP assay is cost-effective and easily implementable for routine clinical monitoring of CRC recurrence, which can lead to better patient management after surgery.
Plasma DNA extraction was performed using 2 to 5 mL of plasma with the Apostle MiniMax High-Efficiency cfDNA Isolation Kit, according to the product manual.
Cell-free chromatin immunoprecipitation can determine tumor gene expression in lung cancer patients. Christoffer Trier Maansson, Peter Meldgaard, Magnus Stougaard, Anders Lade Nielsen, Boe Sandahl Sorensen. Molecular Oncology. February 24, 2023. DOI: 10.1002/1878-0261.13394.
(Note: Apostle MiniMax technology is used in this study.)
Cell-free DNA (cfDNA) in blood plasma can be bound to nucleosomes that contain post-translational modifications representing the epigenetic profile of the cell of origin. This includes histone H3 lysine 36 trimethylation (H3K36me3), a marker of active transcription. We hypothesized that cell-free chromatin immunoprecipitation (cfChIP) of H3K36me3-modified nucleosomes present in blood plasma can delineate tumor gene expression levels. H3K36me3 cfChIP followed by targeted NGS (cfChIP-seq) was performed on blood plasma samples from non-small cell lung cancer patients (NSCLC, n = 8), small cell lung cancer patients (SCLC, n = 4) and healthy controls (n = 4). H3K36me3 cfChIP-seq demonstrated increased enrichment of mutated alleles compared to normal alleles in plasma from patients with known somatic cancer mutations. Additionally, genes identified to be differentially expressed in SCLC and NSCLC tumors had concordant H3K36me3 cfChIP enrichment profiles in NSCLC (sensitivity = 0.80) and SCLC blood plasma (sensitivity = 0.86). Findings here expand the utility of cfDNA in liquid biopsies to characterize treatment resistance, cancer subtyping, and disease progression.
(Material and Methods section) -Both the input as well as the cfChIP samples were purified using Apostle MiniMax High Efficiency cfDNA Isolation Kit (Beckman Coulter, Indianapolis, IN, USA) according to manufacturer’s instructions.
Response prediction and risk stratification of patients with rectal cancer after neoadjuvant therapy through an analysis of circulating tumour DNA.
Liu W, Li Y, et al. EBioMedicine March 17, 2022; https://www.thelancet.com/journals/ebiom/article/PIIS2352-3964(22)00129-3/fulltext
Background - Multiple approaches based on cell-free DNA (cfDNA) have been applied to detect minimal residual disease (MRD) and to predict prognosis or recurrence. However, a comparison of the approaches used in different cohorts and studies is difficult. We aimed to compare multiple approaches for MRD analysis after neoadjuvant therapy (NAT) in patients with locally advanced rectal cancer (LARC).
Methods - Sixty patients with LARC from a multicentre, phase II/III randomized trial were included, with tissue and blood samples collected. For each cfDNA sample, we profiled MRD using 3 approaches: personalized assay targeting tumour-informed mutations, universal panel of genes frequently mutated in colorectal cancer (CRC), and low depth sequencing for copy number alterations (CNAs).
Findings - Positive MRD based on post-NAT personalized assay was significantly associated with an increased risk of recurrence (HR = 27.38; log-rank P < 0.0001). MRD analysis based on universal panel (HR = 5.18; log-rank P = 0.00086) and CNAs analysis (HR = 9.24; log-rank P = 0.00017) showed a compromised performance in predicting recurrence. Both the personalized assay and universal panel showed complementary pattern to CNAs analysis in detecting cases with recurrence and the combination of the two types of biomarkers may lead to better performance.
Interpretation - The combination of mutation profiling and CNA profiling can improve the detection of MRD, which may help optimize the treatment strategies for patients with LARC.
(Methods Section) cfDNA was extracted from 1.5-4.5 mL of plasma with the Apostle MiniMax cfDNA isolation kit (C40605, Apostle; San Jose, CA, USA).
A method for early diagnosis of lung cancer from tumor originated DNA fragments using plasma cfDNA methylome and fragmentome profiles.
Yeo Jin Kim, Hahyeon Jeona, Sungwon Jeon, et al. Molecular and Cellular Probes Volume 66, December 2022
Abstract Early detection is critical for minimizing mortality from cancer. Plasma cell-free DNA (cfDNA) contains the signatures of tumor DNA, allowing us to quantify the signature and diagnose early-stage tumors. Here, we report a novel tumor fragment quantification method, TOF (Tumor Originated Fragment) for the diagnosis of lung cancer by quantifying and analyzing both the plasma cfDNA methylation patterns and fragmentomic signatures. TOF utilizes the amount of ctDNA predicted from the methylation density information of each cfDNA read mapped on 6243 lung-tumor-specific CpG markers. The 6243 tumor-specific markers were derived from lung tumor tissues by comparing them with corresponding normal tissues and healthy blood from public methylation data. TOF also utilizes two cfDNA fragmentomic signatures: 1) the short fragment ratio, and 2) the 5′ end-motif profile. We used 298 plasma samples to analyze cfDNA signatures using enzymatic methyl-sequencing data from 201 lung cancer patients and 97 healthy controls. The TOF score showed 0.98 of the area under the curve in correctly classifying lung cancer from normal samples. The TOF score resolution was high enough to clearly differentiate even the early-stage non-small cell lung cancer patients from the healthy controls. The same was true for small cell lung cancer patients.
(Methods Section) Cell-free DNA was extracted from 3 to 5 ml of plasma using XXX method or Apostle MiniMax High Efficiency Isolation Kit (Beckman Coulter Life Sciences, C40603), according to the manufacturers’ procedures.
High-resolution DNA size enrichment using a magnetic nano-platform and application in non-invasive prenatal testing.
Zhang et al. Analyst. July 2020, 145, 5733-5739 (PDF)
Precise DNA sizing can boost sequencing efficiency, reduce cost, improve data quality, and even allow sequencing of low-input samples, while current pervasive DNA sizing approaches are incapable of differentiating DNA fragments under 200 bp with high resolution (<20 bp). In non-invasive prenatal testing (NIPT), the size distribution of cell-free fetal DNA in maternal plasma (main peak at 143 bp) is significantly different from that of maternal cell-free DNA (main peak at 166 bp). The current pervasive workflow of NIPT and DNA sizing is unable to take advantage of this 20 bp difference, resulting in sample rejection, test inaccuracy, and restricted clinical utility. Here we report a simple, automatable, high-resolution DNA size enrichment workflow, named MiniEnrich, on a magnetic nano-platform to exploit this 20 bp size difference and to enrich fetal DNA fragments from maternal blood. Two types of magnetic nanoparticles were developed, with one able to filter high-molecular-weight DNA with high resolution and the other able to recover the remaining DNA fragments under the size threshold of interest with >95% yield. Using this method, the average fetal fraction was increased from 13% to 20% after the enrichment, as measured by plasma DNA sequencing. This approach provides a new tool for high-resolution DNA size enrichment under 200 bp, which may improve NIPT accuracy by rescuing rejected non-reportable clinical samples, and enable NIPT earlier in pregnancy. It also has the potential to improve non-invasive screening for fetal monogenic disorders, differentiate tumor-related DNA in liquid biopsy and find more applications in autoimmune disease diagnosis.
Detecting SARS-CoV-2 variants in wastewater and their correlation with circulating variants in the communities. Lin Li, Timsy Uppal, Paul D. Hartley, Andrew Gorzalski, Mark Pandori, Michael A. Picker, Subhash C. Verma & Krishna Pagilla. Scientific Reports volume 12, Article number: 16141 (2022) , September 2022
Abstract
Detection of SARS-CoV-2 viral load in wastewater has been highly informative in estimating the approximate number of infected individuals in the surrounding communities. Recent developments in wastewater monitoring to determine community prevalence of COVID-19 further extends into identifying SARS-CoV-2 variants, including those being monitored for having enhanced transmissibility. We sequenced genomic RNA derived from wastewater to determine the variants of coronaviruses circulating in the communities. Wastewater samples were collected from Truckee Meadows Water Reclamation Facility (TMWRF) from November 2020 to June 2021. SARS-CoV-2 variants resulting from wastewater were compared with the variants detected in infected individuals' clinical specimens (nasal/nasopharyngeal swabs) during the same period and found conclusively in agreement. Therefore, wastewater monitoring for SARS-CoV-2 variants in the community is a feasible strategy as a complementary tool to clinical specimen testing in the latter's absence.
(Methods Section) The NP swabs received at the Nevada State Public Health Laboratory (NSPHL) from the Reno-Sparks metropolitan for SARS-CoV-2 diagnostic testing were subjected for RNA extraction using Virus Total NA Isolation Fast Kit (Apostle MiniGenomics, San Jose, CA, USA), followed by RT-PCR for the detection of SARS-CoV-2 genome using US Federal Drug Administration Emergency Use Authorization (FDA-EUA) diagnostic kits, as described previously.
Rapid repeat infection of SARS-CoV-2 by two highly distinct delta-lineage viruses.
Andrew J. Gorzalski , Christina Boyles, Victoria Sepcic, Subhash Verma, Joel Sevinsky, Kevin Libuit, Stephanie Van Hooser, Mark W. Pandori. Diagnostic Microbiology and Infectious Disease. Volume 104, Issue 1, September 2022, 115747; https://doi.org/10.1016/j.diagmicrobio.2022.115747
An instance of sequential infection of an individual with, firstly, the Delta variant and secondly a Delta-sub-lineage has been identified. The individual was found positive for the AY.26 lineage 22 days after being found positive for the Delta [B.1.617.2] variant. The viruses associated with the cases showed dramatic genomic difference, including 31 changes that resulted in deletions or amino acid substitutions. Seven of these differences were observed in the Spike protein. The patient in question was between 30 and 35 years old and had no underlying health conditions. Though singular, this case illustrates the possibility that infection with the Delta variant may not itself be fully protective against a population of SARS-CoV-2 variants that are becoming increasingly diverse.
Nucleic acid extractions were performed by Apostle MagTouch Nucleic Acid Extraction Automation Systems [Apostle Inc, San Jose, CA].
Combining cell-free RNA with cell-free DNA in liquid biopsy for hematologic and solid tumors. Maher Albitar, Hong Zhang, Ahmad Charifa, et al. Heliyon 9 (2023) e16261; May 16, 2023
(Note: Apostle MiniMax technology is used in this study.)
Abstract Current use of liquid biopsy is based on cell-free DNA (cfDNA) and the evaluation of mutations or methylation pattern. However, expressed RNA can capture mutations, changes in expression levels due to methylation, and provide information on cell of origin, growth, and proliferation status. We developed an approach to isolate cell-free total nucleic acid (cfDNA) and used targeted next generation sequencing to sequence cell-free RNA (cfRNA) and cfDNA as new approach in liquid biopsy. We demonstrate that cfRNA is overall more sensitive than cfDNA in detecting mutations. We show that cfRNA is reliable in detecting fusion genes and cfDNA is reliable in detecting chromosomal gains and losses. cfRNA levels of various solid tumor biomarkers were significantly higher (P < 0.0001) in samples from solid tumors as compared with normal control. Similarly, cfRNA lymphoid markers and cfRNA myeloid markers were all higher in lymphoid and myeloid neoplasms, respectively as compared with control (P < 0.0001). Using machine learning we demonstrate cfRNA was highly predictive of diagnosis (AUC >0.98) of solid tumors, B-cell lymphoid neoplasms, T-cell lymphoid neoplasms, and myeloid neoplasms. In evaluating the host immune system, cfRNA CD4:CD8B and CD3D:CD19 ratios in normal controls were as expected (median: 5.92 and 6.87, respectively) and were significantly lower in solid tumors (P < 0.0002). This data suggests that liquid biopsy combining analysis of cfRNA with cfDNA is practical and may provide helpful information in predicting genomic abnormalities, diagnosis of neoplasms and evaluating both the tumor biology and the host response.
(Methods section) We used Apostle MiniMax High Efficiency cfRNA/cfDNA isolation kit and followed the recommended protocol. After extraction, half of the cfDNA was treated with DNase to obtain cfRNA and the other half was used for DNA studies.
For a complete list of publications citing Apostle technologies, see Publications.